Learning to Culture the Uncultured
Cultivation as a magic-free learning system
Imagine that tomorrow we sequence every living thing on Earth. Every pond, biofilm, sediment layer, and obscure microbial eukaryote hiding in environmental DNA enters the databases. Phylogenies bloom. New branches appear everywhere.
And then, after the initial thrill, we discover that most of this new biology is still behind glass. Named, placed on a tree, and partially inferred from sequence, but not experimentally available. We cannot perturb it, rear it, starve it, rescue it, watch it recover, or expose it to a new environmental rhythm. It exists as a sequence-shaped ghost.
The loop in its most literal form: find the organism, bring it into a controlled world, and keep changing the world until it stops merely surviving and starts living.
This is the culturomics bottleneck, and it is stranger than a missing recipe.
The obvious problem is that we do not know the right medium, temperature, pH, partner organism, oxygen level, light cycle, nutrient pulse, or stress regime. But that is incomplete. The deeper problem is that cultivation is treated as recipe discovery, when it is often closer to rearing a living, sensing, history-dependent system through a landscape the cultivator does not fully understand either. In fact, we probably have way more unknown unknowns than we care to admit.
The deeper failure is that culturing knowledge does not accumulate well. It leaks. It lives in people, in half-remembered tricks, in incubator habits, in the one postdoc who knew which bacterial lawn looked right. Success becomes a method section. Failure becomes ether. A dead culture rarely becomes a data point someone else can use.
This is not because the people doing the work are insufficiently clever. It is because the incentives point elsewhere. The scientific system rewards discoveries made after an organism is cultured, not the long, careful, negative, ugly work required to culture it. A conventional grant can buy a better attempt inside one lab. It rarely builds the maintained machinery that lets failure become shared evidence.
The bet is simple and dangerous enough to matter: many difficult cultures fail not because we missed the right recipe, but because we erased the history that would have made the recipe work. The organism was not brought through the right sequence of cues, recoveries, stresses, partners, and absences. We keep asking what condition it likes. Sometimes the better question is what it had to live through before that condition became survivable.
This is not hypothetical for me. In Capsaspora, we recently showed predictive cue-stress pairing reduced mortality from 26.8% to 12.1% relative to non-predictive cue timing. The number matters, but the insult it delivers to the recipe view matters more. The organism was not responding only to a condition. It was responding to a relationship between cue, stress, and time.
What I'm actually trying to avoid here are vibes. Often, a highly skilled, super-intuitive lab-tech will tell you a culture looks healthy, it's doing well! We should try our best to avoid precisely that. The endpoint is time to stable, reproducible culture, is what matters and what we should solve. Stable means something boring and hard: target-lineage persistence across a pre-registered window, for example, 3-5 passages or several weeks, depending on generation time, identity confirmed by marker sequencing, viability above threshold, and reproduction in independent chambers. In mixed enrichments, target abundance and partner-community stability both count; a beautiful bacterial soup that loses the protist is a highly informative failure. Secondary endpoints include survival after stress, recovery dynamics, morphology, model calibration, and the number of physical experiments saved.
The first version should be ugly, dirt cheap, easy enough to build, and be allowed to fail. Build 12-24 low-cost programmable incubators using 3D-printed housings, commodity sensors, microcontrollers, LEDs, temperature control, simple fluid handling, and optional camera modules. A plausible target is roughly $200-$400 per chamber, with an initial hardware budget under $10,000 before shared microscopy and sequencing costs.
The pilot should have two phases. Phase one uses a tractable organism such as Capsaspora owczarzaki. Capsaspora is already culturable, so the question there is not whether the machine can perform a miracle (an often-used term for a successful culture). The question is whether it can predict stress recovery, morphology, survival, and state transitions under controlled cue histories better than static protocols can. Phase two moves to one difficult protist and one mixed-enrichment sample where bacterial partners may matter. That is where true culture establishment is tested.
Each phase should face the boring baselines: expert-designed protocols, random exploration, static optimization, and blind continuous incubation. Same chamber count. Same inoculum. Same observation schedule. Same experimental budget. Same stopping rules. Otherwise, the loop might win simply because it was watched more carefully, which is the wrong kind of victory.
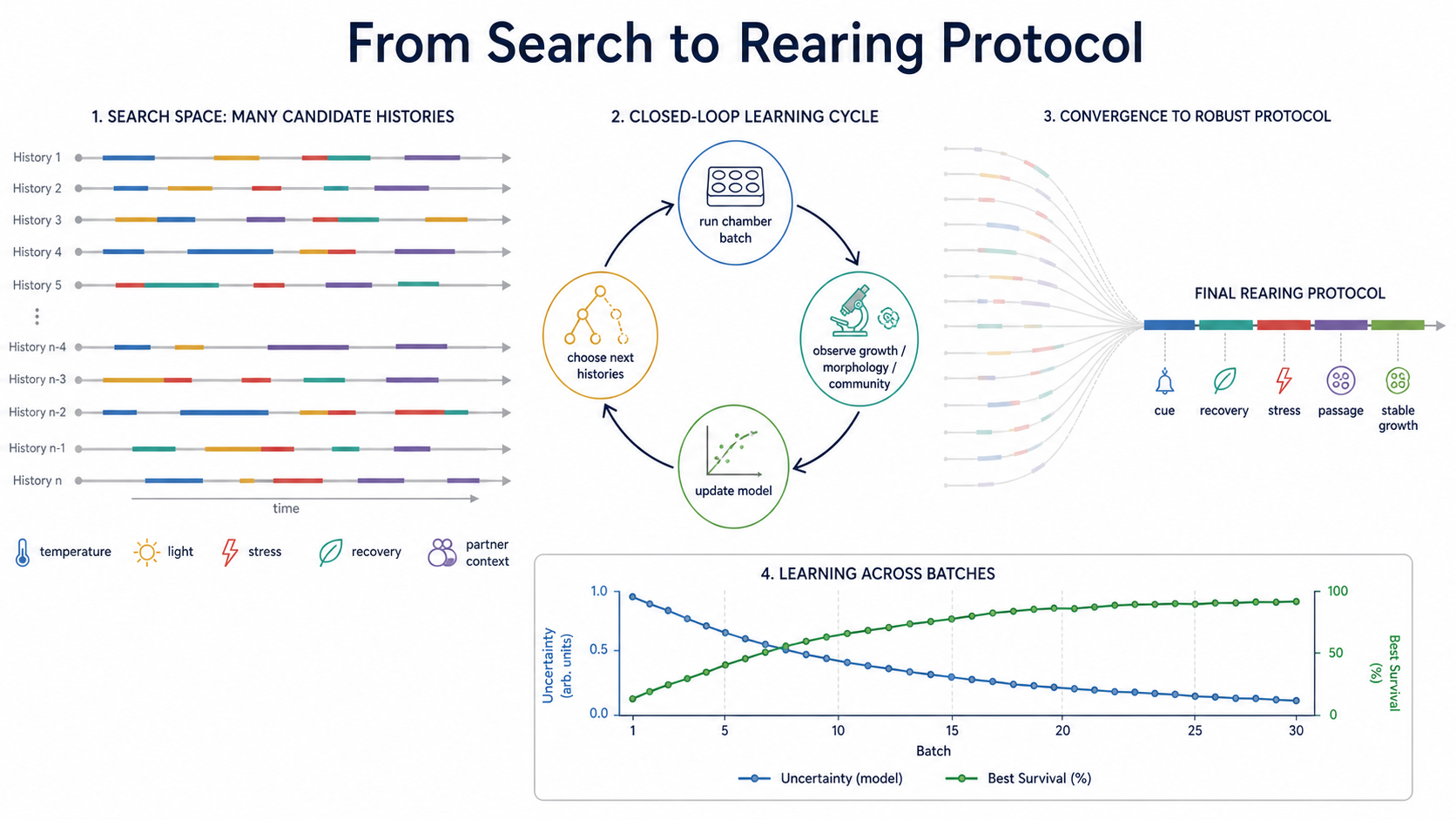
The closed-loop arm should not merely optimize growth conditions. It should optimize rearing schedules: cue reliability, stress timing, recovery intervals, environmental stochasticity, and partner-community context. A static optimizer asks, "Which condition works?" A rearing platform asks, "Which structured history lets the organism become capable of surviving this condition?"
A condition is a point. A rearing protocol is a path
This is where the AI part can easily become stupid. If it merely automates the old screen, it is just a faster way to misunderstand the organism. The useful loop asks a different question: not which condition is best, but which history makes the organism capable of living here.
The loop earns its keep only if it reaches stable culture faster, improves recovery, or achieves equivalent performance with fewer physical experiments than expert, random, blind-continuous, or static-optimization protocols. If it only produces prettier models without improving experimental decisions, it fails. No practical win, no victory. Simple. Traceable. Accountable.
That failure would still matter. The point is not to promise a universal culturing machine. The point is to test whether cultivation can become a learning system.
The timing is not mystical. Three ordinary things have become cheap enough to combine. Low-cost embedded hardware can make controlled environmental schedules replicable. Computer vision can turn microscopy into continuous phenotypic readouts rather than occasional inspection. Active-learning models can treat failed culture attempts as informative data rather than wasted experiments. Together, these parts can turn artisanal troubleshooting into a cumulative experimental loop.
A protist is not agar with preferences. It is not a passive object waiting for the correct salt concentration. Its future viability depends on sensing, state transitions, preparation, and recovery. Sometimes it needs a partner, a cue before a stress, the absence of a cue, or a week of structured exposure before Tuesday's lethal condition becomes tolerable.
Current culturomics infrastructure is not built for this, and probably exactly the opposite of it is needed. A culture fails. Someone changes the medium. It fails again. Someone remembers that a related organism liked a bacterial feeder, or darkness, or patience. A few things are tried. Some work. Most disappear into the private sediment of lab memory. When random success happens, the path of failures is quickly forgotten.
Around that small failure sits a bigger one. The relevant expertise is split across domains that do not naturally live together: protist biology, embedded hardware, microscopy, control theory, machine learning, microbial ecology, automation, and data engineering. A conventional lab can host pieces of this stack. It usually cannot professionalize the whole thing. And it's definitely hard to get someone excited enough to fund.
So the bottleneck persists. Environmental sequencing has made microbial eukaryotic diversity visible, but not experimentally available. We have built telescopes into the hidden biosphere and very few doors.
In the actual box, nothing magical happens. Each chamber runs controlled environmental schedules: temperature, light, oxygen, nutrient pulses, stress timing, recovery intervals, cue reliability, and community context. The system watches through microscopy, growth readouts, survival assays, and, when needed, marker-gene or metagenomic profiling. Every experiment updates a shared state model, and the model chooses the next experiment.
This is not ordinary reinforcement learning dropped onto biology. In a computer game, the world is already written. Cultivation is meaner than that. The organism changes as it is being measured. Failed experiments are not just bad rewards; they mark the boundary of viability, the cost of transitions, and the difference between a dead end and a necessary preparation step. The best version of the platform is a rearing loop: infer, perturb, observe, update, and only then intervene again.
This is not just a metaphor. My own work keeps circling the same wall from different sides. Identification in novel niches can be fragile. Bulk measurements can hide lineage-level fates. Exposure history can change the adaptation regime. Sensing and plasticity only pay when environments contain learnable structure. The Capsaspora result is the cleanest version of the pattern: a living system doing better not because the condition was nicer, but because the history made more sense.
Together, these results point to the same missing infrastructure. Culturing systems must identify what is present, control the history and predictability of exposure, and track which subpopulations survive. Identification, rearing history, and lineage-aware population dynamics are not separate problems. They are three parts of one bottleneck.
Capsaspora is a good launch system because it is biologically interesting, experimentally tractable, and surrounded by unusually rich community resources: genome assemblies, life-stage transcriptomics, proteomics, culture methods, transfection protocols, and emerging genetic tools. These data provide priors. They tell us which states might matter, which phenotypes are measurable, and which viability metrics are not nonsense.
After Capsaspora, the real test begins: what transfers, what breaks, which measurements are predictive, and where does the digital twin fail?
Protists are not special in this respect. They are just where the wound is visible to me. The same problem appears in anaerobes that depend on subtle redox histories, syntrophic bacteria whose viability depends on partners, marine larvae that require timed cues, organoid maturation protocols that depend on developmental order, and plant tissue cultures that respond to hormone sequences rather than single media recipes. These are not identical systems, but they share a structure: the relevant experimental unit is not a condition. It is a history.
The pilot must be allowed to fail. If the model cannot beat budget-matched expert protocols, that tells us something. If inferred state variables do not predict future viability, that tells us something. If gains vanish in independent chambers, that tells us something. A good bottleneck experiment should not require triumph to be informative.
That is why the output cannot be a clever box in one lab. It has to be a public object: chamber bill of materials, CAD files, firmware, calibration assays, metadata schema, image-analysis pipeline, benchmark tasks, protocols, failure taxonomy, and failed-trial records in reusable formats. Especially failed experiments. A dead culture is not just a disappointment; it is a coordinate on the map. Right now, the map is mostly stored in tired people's heads.
This is the institutional object culturomics is missing: an open protocol-learning commons that lets another lab use the accumulated search without inheriting all the original tacit craft. It is not glamorous in the usual academic sense. It is the kind of maintained, negative-result-rich infrastructure that can unlock whole fields.
Cultures turn biodiversity from description into mechanism. They allow genetics, physiology, perturbation, imaging, and eventually biotechnology. They let us ask whether a lineage contains useful biomolecules, antibiotic-relevant functions, climate-relevant physiology, or simply a way of being alive that our current model organisms never discovered. Sequencing tells us the world is full of such organisms. Cultivation decides whether we get to meet them.
Biology has spent centuries learning from the organisms that were easy to grow. This was reasonable. It gave us yeast, flies, worms, mice, E. coli, and a civilization of experimental power. But ease of growth is not the same as importance. It is a sampling bias wearing a lab coat.